Making Scientific Tools Accessible Through Agentic Interfaces
February 15, 2026
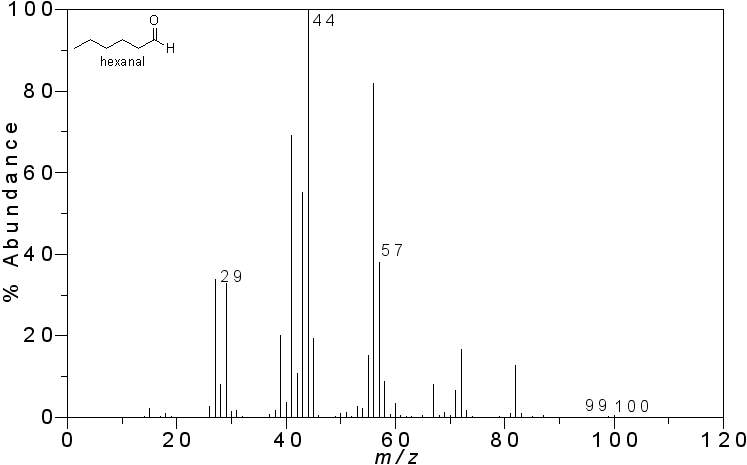
Mass spectrum of hexanal. Image by Vladislav Andriashvili, CC BY-SA 3.0, via Wikimedia Commons.
Scientific computing has a last-mile problem. Researchers write or commission code that solves their analysis task, and then it sits on one person’s laptop behind a CLI or a Jupyter notebook. The people who most need the tool, the ones running the experiments, can’t use it without help from whoever wrote it.
My dad reminded me of this over the holidays. He’d spent the new year identifying molecular formulas from mass spectrometry peaks. Here’s the problem: a mass spectrometer measures the mass-to-charge ratio (m/z) of ionized molecules. You get a number like 1519.154, and you need to figure out which molecular formula produces that exact mass. For his system (yttrium-manganese clusters with tert-butylcarboxylate ligands), that means searching every combination of Y, Mn, tBuCOO, O, H, C, F, and N atoms until something matches within a few parts per million. It’s a brute-force combinatorial search over eight elements, and it’s tedious to do by hand.
I built a CLI tool. It works. But “install Python 3.11, clone the repo, run ./search.py 1519.154 --ppm 5 -c 1” is not something you hand to a chemist (especially my dad, who hasn’t touched a terminal since like 2003). So I wrapped it as an MCP server. Now he asks Claude what formulas match a peak and gets an answer.
The interesting part isn’t the LLM-instead-of-CLI trick. It’s that wrapping the tool this way turns a one-off script into a typed, contract-driven computational primitive that agents can reason about and orchestrate.
The tool
The formula space is combinatorial in nature: there are eight distinct elements, each spanning a defined range of stoichiometric counts. I constructed the full search space using NumPy’s meshgrid to generate multidimensional arrays of all possible combinations, then pruned until all that’s left are the plausible formulas.
y, mn, k, o, h, c, f, n = np.meshgrid(
y_vals, mn_vals, k_vals, o_vals, h_vals, c_vals, f_vals, n_vals,
indexing="ij",
)
y, mn, k, o, h, c, f, n = [a.ravel() for a in (y, mn, k, o, h, c, f, n)]
...
masses = (y*MASS[metal] + mn*MASS["Mn"] + k*MASS["tBuCOO"]
+ o*MASS["O"] + h*MASS["H"] + c*MASS["C"]
+ f*MASS["F"] + n*MASS["N"])
hits = np.abs(masses - target_mass) <= target_mass * ppm * 1e-6It’s completely brute-force, but NumPy does the iteration in C so it’s pretty fast.
Wrapping it for agentic use
The Model Context Protocol lets you expose functions as tools an LLM can call. FastMCP makes the wrapping very minimal:
from mcp.server.fastmcp import FastMCP
from pydantic import BaseModel, Field
mcp = FastMCP("formula_search_mcp")
class FormulaSearchInput(BaseModel):
mz: float = Field(..., description="Observed m/z value", gt=0)
ppm: float = Field(default=10.0, description="PPM tolerance", gt=0, le=100)
mode: IonMode = Field(default=IonMode.NEGATIVE)
coarseness: int = Field(default=2, ge=1, le=3)
metal: Metal = Field(default=Metal.Y)
@mcp.tool(name="formula_search_mz")
async def formula_search_mz(params: FormulaSearchInput) -> str:
hits = _search_formulas(
mz=params.mz, ppm=params.ppm,
mode=params.mode.value, coarseness=params.coarseness,
metal=params.metal.value,
)
return _format_results_markdown(hits)Sensible defaults mean my dad can say “search for 1519.154” without specifying PPM tolerance, ion mode, or coarseness. But if he says “try that again with stricter constraints,” Claude sets coarseness=1 because the field description says 1=strict.
There is also a formula_scan_all_levels utility that executes all three coarseness tiers in a single pass and returns the hits for each level. This enables Claude to determine, in one invocation rather than three, which formulas are clear matches under strict constraints and which emerge only after relaxing those constraints.
Why not just build a GUI?
Streamlit, Gradio, and web forms all work, but they each require building and maintaining a separate interface, and they’re still “an app.” The researcher has to learn where to click, what fields mean, what values are reasonable.
The difference with an agentic interface is that it’s conversational and adaptive. My dad doesn’t need to know the parameters exist. He says “I have a peak at 1519.154 in negative mode, what could it be?” Claude calls the tool, gets a table of candidate formulas with PPM errors, and explains the matches. He says “try lanthanum instead of yttrium” and Claude calls with the correct arguments. He doesn’t navigate a UI. He describes what he wants and gets an answer.
A GUI also only serves humans. An MCP server serves both humans (through an agentic interface) and agents (through direct tool calls). An automated pipeline could call the same tool the same way my dad does through Claude.
Why this matters
People in software engineering take CLI experience for granted. Most people in the world have never seen a console in their lives, and will swiftly run away from your product the first time they see a text-only interface.
This is especially the case in research. There are groups who produce code as part of their research, but the code is barely packaged and often has broken dependencies. In fact, it’s often the case that the researcher who wrote the code themselves won’t be able to decipher it after a few months.
Wrapping an existing tool as an MCP server is not much work. It’s typically only a Pydantic model and a decorator, and often this is doable with coding agents like Claude Code. But the payoff is disproportionate: a one-off script becomes a typed primitive with a schema that agents can discover, reason about, and compose. It becomes usable by anyone who can describe what they want in words.
A lot of custom analysis code gets written once, used by one person, and abandoned when they leave. This is a way to make it stick around.
The repo is at github.com/MatchaOnMuffins/ms-cluster-formula-search.